
In the race to create more potent and stable medicines, scientists know that adding fluorine can improve drug molecules.
However, there is only one fluorination reaction known to occur in nature. And this process is performed by complex and highly specialized enzymes that are difficult to replicate. Now, scientists have developed a new, streamlined molecule that can do the work of enzymes but be designed and controlled minutely. This new catalyst transforms a safe, affordable fluoride salt into fluorinated organic molecules. In addition to improved pharmaceuticals, fluorinated molecules can also act as useful “radiotracers” in high-resolution 3-D medical imaging technology, such as PET (positron emission tomography) scanning.
Robert Paton, an associate professor in the Department of Chemistry at Colorado State University, contributed to the new study, which was published this week in Science. The work was conceived and led by Professor Véronique Gouverneur at the University of Oxford, with design and synthesis provided by Gabriele Pupo and Francesco Ibba in the Gouverneur lab, and the mode of action uncovered using quantum mechanical calculations by Oxford’s David Ascough and Paton.
“This new catalyst performs the same function as the biological enzyme, but is a fraction of the size and is more widely applicable,” Paton said. In addition, the designed catalyst can be used to incorporate fluorine into a variety of organic molecules, whereas the naturally occurring enzyme is limited to just one target.
Designing a new molecular claw
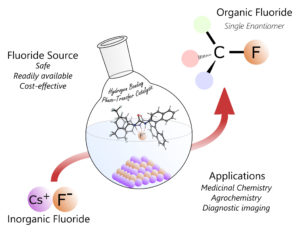 Catalysts speed up chemical reactions, creating new compounds from the original ingredients. The catalyst that Paton and his colleagues developed is specifically tuned to integrate fluorine atoms into a new organic product. “It works like a claw crane arcade game,” Paton said of the catalyst’s mode of action. “You’re trying to pick one fluffy toy out of many and hold on tightly. In a broad sense, that’s what our catalyst does. Surrounded by other chemicals, it selectively binds fluoride anions using three hydrogen-bonds – effectively a molecular claw – and then positions it to add the substrate,” or target location.
Catalysts speed up chemical reactions, creating new compounds from the original ingredients. The catalyst that Paton and his colleagues developed is specifically tuned to integrate fluorine atoms into a new organic product. “It works like a claw crane arcade game,” Paton said of the catalyst’s mode of action. “You’re trying to pick one fluffy toy out of many and hold on tightly. In a broad sense, that’s what our catalyst does. Surrounded by other chemicals, it selectively binds fluoride anions using three hydrogen-bonds – effectively a molecular claw – and then positions it to add the substrate,” or target location.
With such specific tasks, catalysts are extremely challenging to create from scratch. “You’re trying to construct a molecule with a specific function and a precise three-dimensional shape,” Paton said.
That’s where Paton’s work comes in. His lab specializes in computational analysis and uses quantum mechanics to predict structure and behavior. In creating a new catalyst, “computer-aided design allows you to visualize atomic-scale structures – in the same way you would design a new airplane or new house,” he said. In the past, he added, “the discovery of new functional molecules has relied heavily on serendipity and trial and error. We’ve combined computation with experiment to understand the mechanism of how this thing works. And we applied that early on to guide our experimental design.”
This approach allows them to rely less on time-consuming trial and error. Instead, “it’s about minimizing failure through a better understanding of how reactions take place,” he said.
A greener chemistry
 Paton joined the CSU College of Natural Sciences this January from his post as an associate professor at Oxford University in the U.K., where the new work was conducted. He brings his broadly applicable research to the new Chemistry Research Building.
Paton joined the CSU College of Natural Sciences this January from his post as an associate professor at Oxford University in the U.K., where the new work was conducted. He brings his broadly applicable research to the new Chemistry Research Building.
Beyond the work on fluorine, his research overall is also helping to create a “greener” chemistry. The computational approach cuts down on trial-and-error attempts. And improving catalysis with smarter, custom-created molecules also cuts down on waste products.
Novel catalysts could also be put to work generating new, useful products out of waste. For example, specifically designed compounds could turn discarded plant products into useful chemicals, such as energy products. It is about “being able to control catalysis to generate the products you want,” Paton said. And that work might someday be behind a better pharmaceutical or cleaner fuel – all thanks to a more efficient molecular claw.