Movement signals life, and nowhere is this truer than inside a living cell. The millions of proteins and molecules within each of our cells bend, travel and conform in a complex but orchestrated pattern, regulated by the genes that encode what goes where and when. As part of that pattern, an important class of proteins called dynein transport and deliver various cellular cargoes between different areas of the cell.
Colorado State University biochemistry researcher Steven Markus is particularly intrigued by these large, intracellular motor proteins that move methodically along a network of filamentous tracks called microtubules.
How important is dynein? If dynein were to disappear, we wouldn’t live past a few mitotic cellular divisions. And many neurological diseases, including one called lissencephaly, are linked to defects in dynein function. The goal of many labs, including Markus’, is to understand why.
His research team has made a leap in that understanding by unveiling, in intricate detail, the mechanism by which one particular molecule affects dynein function. While it was long known that the lissencephaly-1 gene, or Lis1, affects dynein activity, the details were unclear. Markus and his team have revealed exactly how Lis1 activates dynein by preventing dynein’s ability to turn itself off, stabilizing it in an “open,” uninhibited conformation.
The new finding flies in the face of previously accepted views that Lis1 acted as an inhibitor of dynein. According to the Markus lab’s new study, published April 27 in Nature Cell Biology, the exact opposite is true: Lis1 activates dynein, working to wedge itself in such a way that the motor protein is prevented from folding itself into an “off” state – inhibiting its ability to auto-inhibit, the researchers explain.
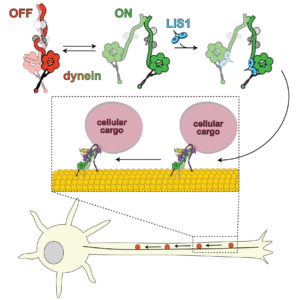
The lissencephaly-1, or Lis1 protein, activates the dynein motor so it can transport cellular cargo. The dynein switches between “off” (left) and “on” (right). Lis1 binds to dynein when it is on, preventing the dynein from switching to an off state.
Understanding molecular basis of disease
A person with lissencephaly, or “smooth brain,” suffers seizures and limited motor function and rarely lives past a few years of age. This devastating disease is associated with a mutation in Lis1, a gene that encodes a critical regulator of dynein.
“I’m interested in the molecular basis for these diseases,” said Markus, assistant professor in the Department of Biochemistry and Molecular Biology. “There will be no therapeutic interventions without understanding how these molecules function.” Beyond that, Markus says, “molecular motors are fun, because we can purify these motors and watch them walk on microtubules in real time using fluorescence microscopy” – which is exactly what the team did for their study.
To carry out their experiments, the researchers employed budding yeast cells as a model system. In contrast to human cells in which dynein performs numerous activities, dynein only performs a single function in these cells. Their findings with this simplified system can translate into human and other higher eukaryotic cells, in which basic dynein function is preserved throughout millions of years of evolution.
The researchers employed several techniques to make their conclusions. The most important was real-time single-molecule imaging. Using a high-yield technique they developed in the lab, the team purified dynein, added a fluorescent molecule, and assembled microscope imaging chambers with purified microtubules to watch the dynein “zip along,” Markus said. This technique allowed them to establish the role of the auto-inhibited conformation in dynein motility.
They also used electron microscopy to take very high-resolution still pictures to determine if the dynein molecules indeed adopted an auto-inhibited conformation, which was unclear when they began their study. “On day one at the electron microscopy facility, we were very surprised to see that the dynein molecules were unambiguously in an auto-inhibited conformation,” Markus said. “It had that very distinct shape.”
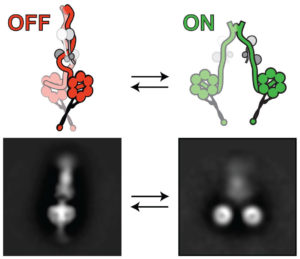
The researchers used a combination of cutting-edge techniques to draw their conclusions, including high-resolution electron microscopy. They used this to visualize the dynein motor in its “off” (left) and “on” (right) states.
The study’s lead author was former graduate student Matthew Marzo (now a postdoctoral researcher at Columbia University), who designed and led the experiments, with assistance from co-author and then-undergraduate student Jacqueline Griswold (now a Ph.D. student and NSF graduate research fellow at Johns Hopkins School of Medicine).
Markus plans to conduct other experiments, using the same yeast cells, to further probe the role of Lis1 in what he and colleagues think is a multi-step pathway that activates dynein. He also hopes to work with neuroscientists at CSU to determine whether the Lis1 activation mechanism functions similarly in neurons. There, the goal will be to provide even further knowledge into how brain diseases like lissencephaly occur at the molecular level.